AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Western blot quantification using imagej4/16/2024  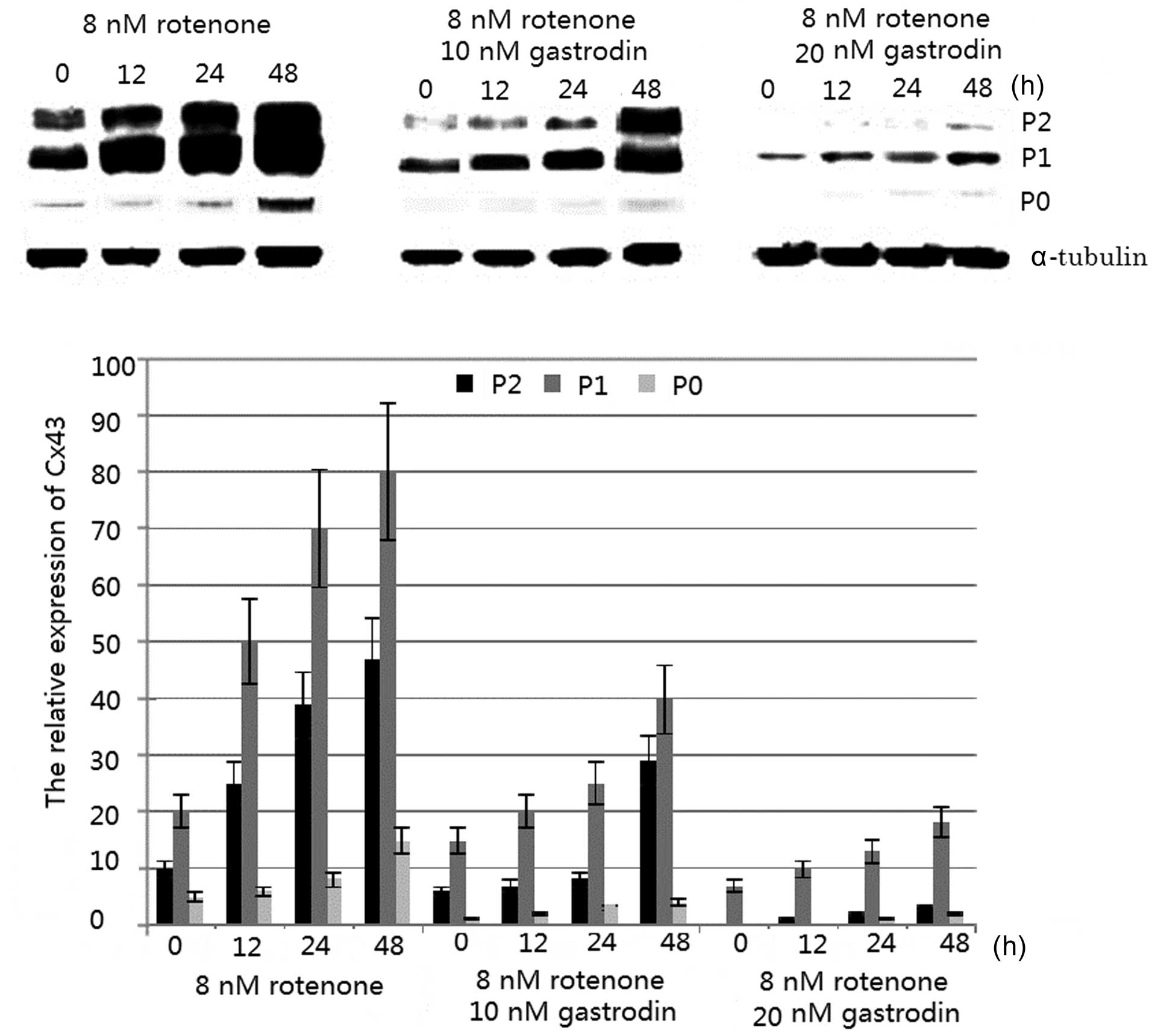
Actually one member of our lab once tried the loading control, where he loaded the same protein lysates at scaler amounts, say, 2ug, 4ug, 8ug,20ug. There are multiple choices outside and what we use in our lab is traditional film system and it's not so sensitive to distinguish slight difference. 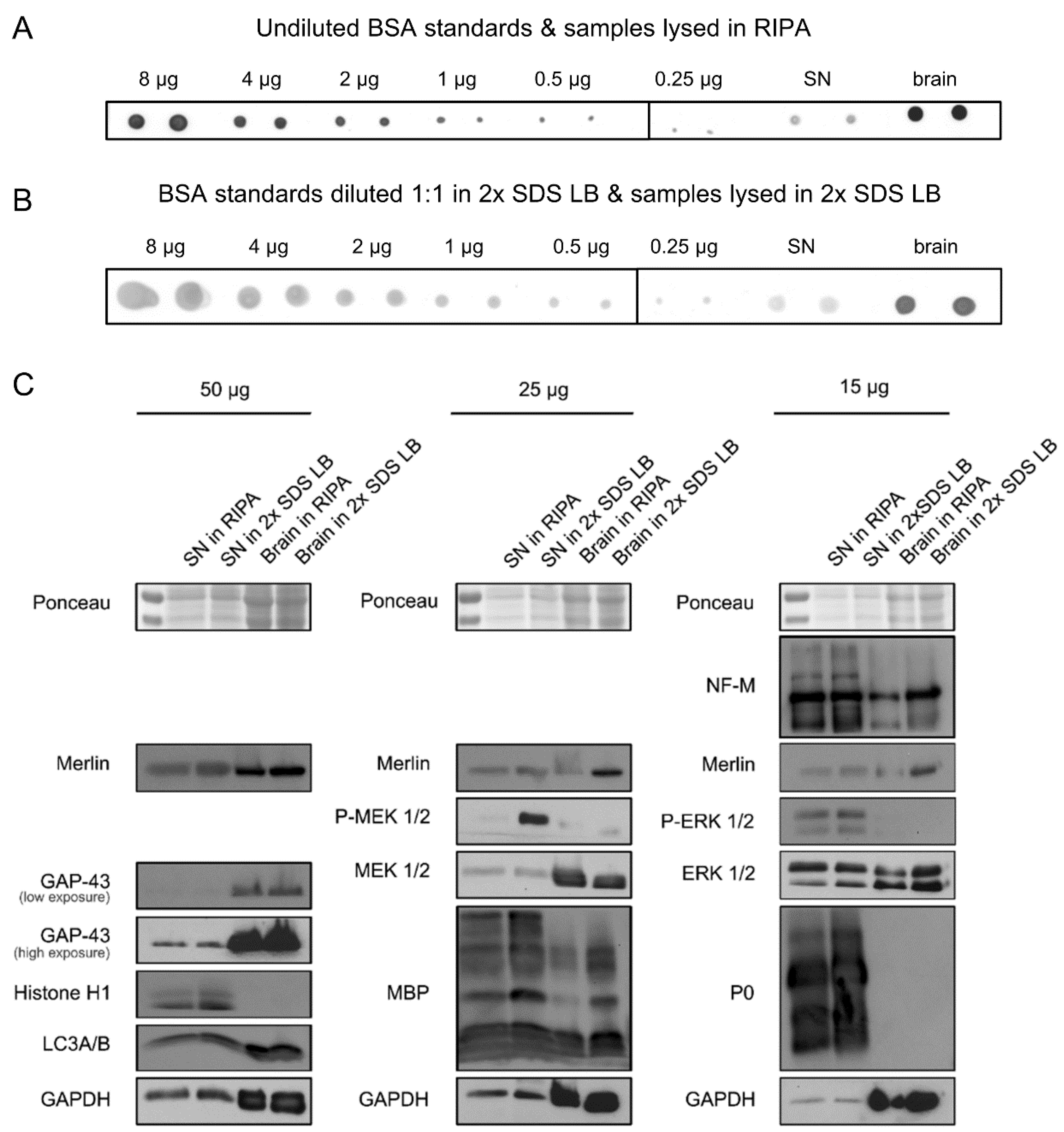
First thing is the sensitivity of your developing reagent and films or the imaging system you use. There are a couple of factors that could make ImageJ analysis tricky. This doesn't mean it's not helpful at all but I would recommend use ImageJ to analyze band intensity with caution. Actually I've done many blots in past few years, but only a few times when I really used the ImageJ and wanted to see the differences in intensity. This is really an interesting post to me as I've been wondering this for quite a long time ago when I first did Western Blot and was told that I could use the ImageJ to analyze the bands. Like Amanda mentioned all of the quantification goes back to the significance and the magical p<0.05! However, seeing is believing so when we see that your lane 1 has a thinner band when compared to lane 6 can we take that as experimentally relevant? Or will we always be trapped in the paradigm of significance. Even though differences in the amount of protein can sometimes be detected with the naked eye quantification with the software we must be aware of the bias we introduce. Even though the lanes are parallel once you take the picture you have to take into account the noise, the amount of protein loaded into your gel and as you mentioned the bias created when you are responsible for selecting the correct area for the quantification. We recently had to perform a couple (always trying to please grant reviewers) and while I was analyzing the data using ImajeJ I started to wonder the exact same thing. I find this post extremely interesting, my lab is a physiology lab and we try not to do western blots unless we absolutely have to. Since your analysis says there is, it is important to know if this difference is biologically relevant. Again, this is all just me observing this band without knowing the scientific relevance (or really the experiment at all), but when I look at those bands I do not see a difference in intensities. This also brings to light a point that sometimes we rely to heavily on the "statistical significance" of these band intensities. Or if it would just be another layer of area selection bias. Me wonder if the loading control would help. Regardless your analysis could shows the consequences of these types of analyses, and it makes I realize this was just a quick experiment of intrigue, and it looks like maybe that top band is the control, or a spurious band. I am curious if you did the same analysis with a loading control, and then normalized the signal. Something like a technical replicate of the area selection? As you mentioned, it is important to correct for noise and that is another added step of variance and potential bias. It makes sense that there is variation between your attempts because of the area around the band, and I wonder if it would be possible to take that into consideration when you are doing western blot analysis. Any help will be appreciated.This is a really interesting analysis, that I have always wondered about. I really just want the raw values for each individual band that corresponds to what I see visually on my blots. It may be just how Image J works so does anyone know how to override this function. I am new to Image J and I am wondering why Image J relativizes the lighter bands when darker bands are measured together. This is strange since you would expect Band C’s intensity be the same or similar to that of Band A. When I measure Lane 2 with Band B and Band C, Band C has a much smaller intensity value, lets say 3000, than Band A. Band C has the same darkness as Band A visually and Band B is darker than Band C. In lane two, there are two bands, Band B and Band C. Now lets say that I am measuring another lane, Lane 2. Band A is a light band and when I measure the intensity value of Band A, Iets say I get 6000. I have only one band, Band A, in Lane 1 of my gel that I am imaging. So when I measure different bands at the same time using the rectangle tool, the lighter band intensity values always seem to be scaled to the darkest band selected. I am having trouble measuring Western blots. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
